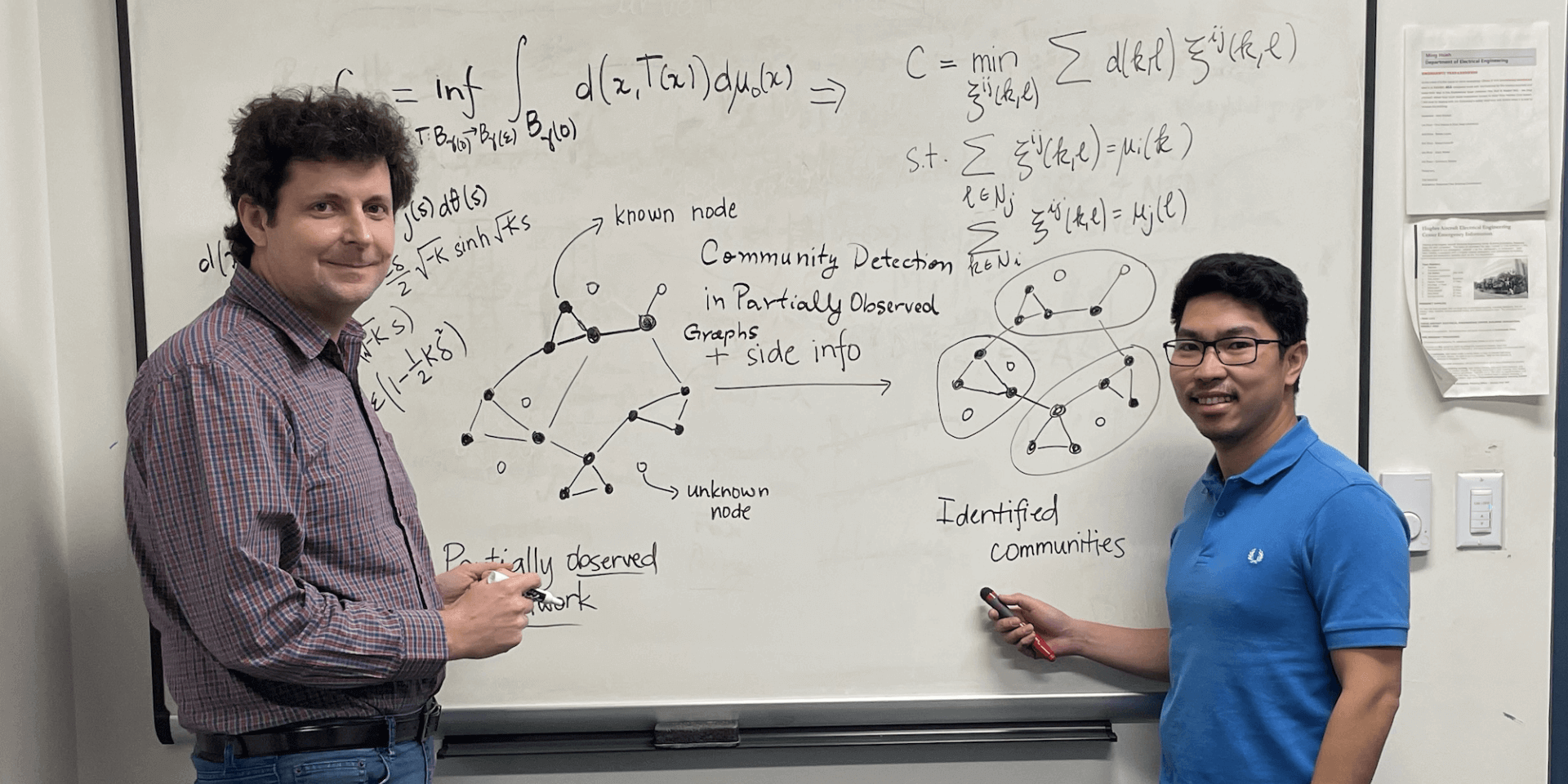
Paul Bogdan (left) and PhD student Jayson Sia (PHOTO CREDIT: USC Viterbi)
Arabidopsis is a small, mostly forgettable weed. But this humble plant is actually one of the most important species, weed or otherwise, anywhere on the planet. It’s what is known as a “model organism” – species that are highly studied by scientists to better understand nature, biology, and even humans. In fact, Arabidopsis is one of the most studied species on earth. Now, data collected from Arabidopsis is the basis for new research from Paul Bogdan, associate professor of electrical and computer engineering at USC Viterbi, his PhD student, Jayson Sia published in Nature, Scientific Reports.
Bogdan and his research group, among other things, specialize in highly complex mathematical models to better understand data represented in graph form. And visualizing complex data in graph form is hugely important. If done correctly, which is no small feat, researchers can analyze these graphs to better understand everything from drug interactions, to radicalization online, to information about genetically engineered plants (more on that last example later).
“One way to make sense of data is to represent it in a graph form. Then, even if we do not know the patterns and rules behind this data, we can try to decipher them by understanding how networks, communities, and other topological varieties might change over time,” Bogdan says.
Today we have more data than ever before. These data sets are the cornerstone for technologies like Machine Learning and AI that make the modern world run. Without being able to quickly access and analyze huge amounts of information, the world we know – and the future engineers are helping to build – could not exist. In other words, without a faster way to make sense of all the information we collect, that bright, shiny, technological future full of self-driving cars and virtual reality and personalized healthcare will never come to fruition. Think of the mathematical models that Bogdan and Sia work on as the engine that powers our future.
So, what does that all have to do with one janky little weed, you might be asking.
What Bogdan and Sia did was take the Arabidopsis protein-protein interaction network and use it as the data set for their mathematical models and graphs. “Arabidopsis is so well studied, and we already have its entire genome sequenced. The scientific community also has a huge amount of data on this plant, which makes it a great model for our research,” said Sia.
And they did this to help solve a huge problem in the graph-making world called “community detection.” Individual data points, or nodes, can be misrepresented on graphs. In fact, they’re often misrepresented. Let’s say you’ve put data from a social network into a graph. Each individual user would be one node on the graph. As the users interact with each other you could, presumably, learn more about how the social network was working. You could even understand how it was evolving and better track things like radicalization online. But if you’re not sure the nodes on your graph are properly represented, you can’t do any of that.
Bogdan and Sia could have chosen any number of graph models to test their theory on. But given the immense impact climate change is already having on the world, they chose to focus on a plant genome so we might better understand how to address food production and sustainability in a changing environment.
“Essentially, we designed a novel mathematical model using the Arabidopsis protein interaction as our map,” said Sia. “Our model not only bypasses the extremely slow process of data analysis and experimental validation, but it also set us on a course to better understanding plant robustness.”
And a better understanding of what makes certain plants stronger is going to be an essential piece of knowledge as climate change continues to wreak havoc across the globe.
Food production is already under threat from climate change in several ways. Not only do changing temperatures make many regions unable to produce food, but deadly plant pathogens and parasites are also moving into new areas faster than ever before. Bogdan and Sia are now using the model they based on Arabidopsis and applying it to other plant species that are resistant to certain pathogens. “We may one day be able to use our model to identify what makes certain species stronger than others. And that could help us engineer new crops that can better survive in a rapidly changing world,” Bogdan said.
This research was in collaboration with Edmond A Jonckheere, professor of electrical and computer engineering at USC Viterbi, David Cook, associate professor of plant pathology at Kansas State University, and Wei Zhang, academic coordinator for the Genomics Core Institute for Integrative Genome Biology at the department of botany and plant sciences at UC Riverside.
Published on November 3rd, 2022
Last updated on November 3rd, 2022












